Project A6
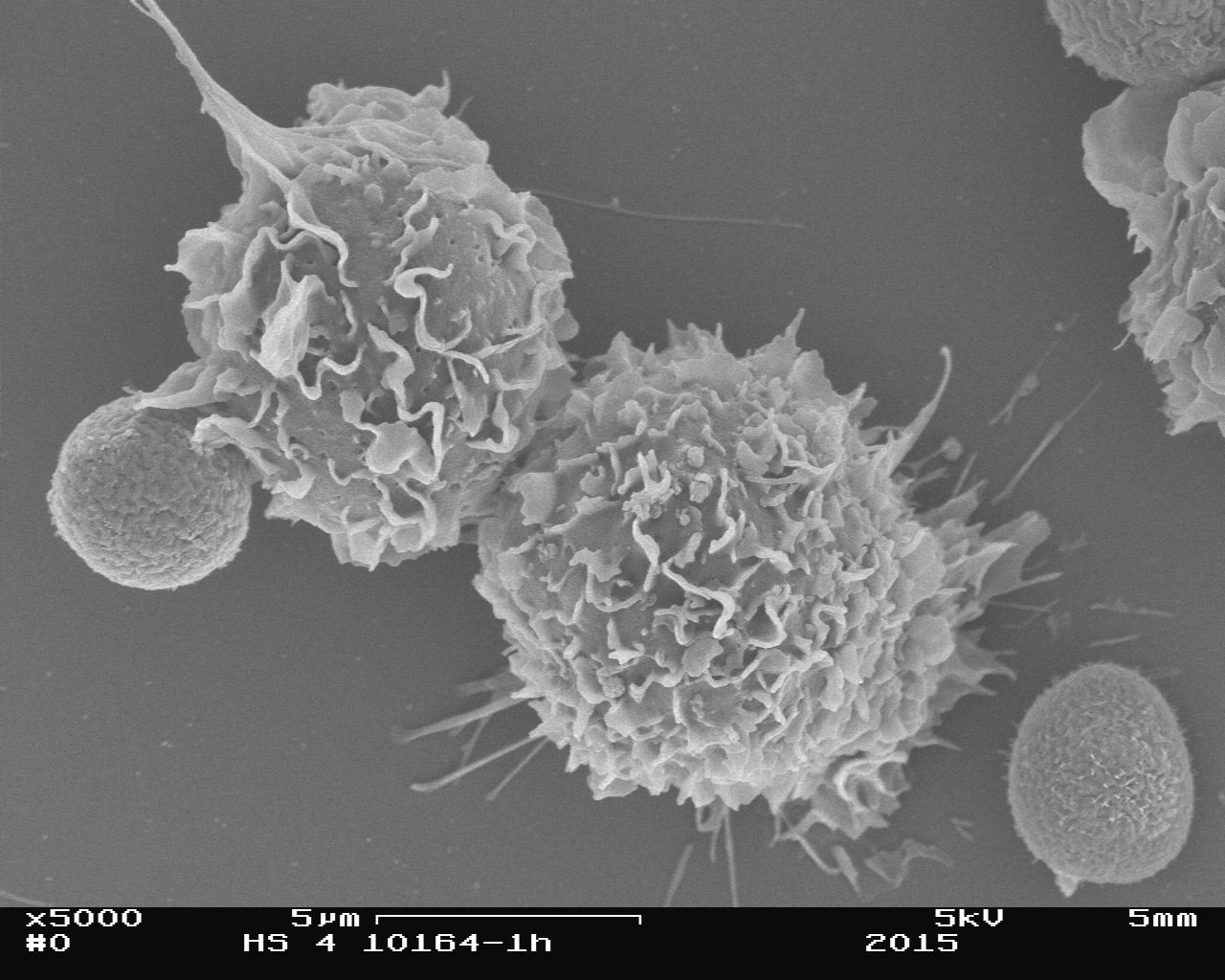
Surface proteins of Lichtheimia corymbifera as virulence determinants
Like Aspergillus fumigatus, Lichtheimia corymbifera is a ubiquitous saprophyte which can cause life-threatening infections, mainly in immunocompromised patients. While these infections (mucormycoses) are uncommon infections, they have been increasingly recognized during the last decades and are associated with high mortality rates. However, only little is known about the pathogenicity mechanisms of these fungi and therapeutic options are still limited.
The cell wall represents the outer border of fungal cells and is crucial for the stability of fungal cells under changing environmental conditions. In fungal pathogens the cell wall also plays a crucial role during the interaction with the host. The main part of the cell wall is composed of carbohydrates such as glucan, chitin and chitosan, which define the interaction with host immune cells. A variety of cell wall located proteins have also been identified in fungi, including adhesins, invasins and hydrolytic enzymes. Such surface proteins are known to contribute to the virulence of fungal pathogens and represent, due to their accessible location, interesting targets for the development of antifungal therapeutics.
In project A6, we aim to get a better understanding of the surface proteins of L. corymbifera and how they are involved in the interaction with the host. Previously, we sequenced and analyzed the genome of L. corymbifera and identified putative target proteins. Using proteomic approaches, we verified the location of several proteins on spore surfaces and identified interesting candidates such as the hydrophobic binding protein HsbA. Since the infection process also involves the hyphal stages of the fungus, stage-specific surface proteomic studies will be conducted. Based on these data we will select target proteins for in-depth molecular and immune biological studies and elucidate their role in the interaction with different host cells.

PD Dr. Kerstin Voigt
Jena Microbial Resource Collection (JMRC)
Department of Microbiology and Molecular Biology
Friedrich Schiller University Jena

Prof. Dr. Axel Brakhage
Project A1 / Project A6 / Project Z1
Institute of Microbiology
Friedrich Schiller University Jena
Department of Molecular and Applied Microbiology HKI
Leibniz Institute for Natural Product Research and Infection Biology - Hans-Knöll-Institute
| Author | Year | Title | Journal | Links |
|---|---|---|---|---|
| Jia LJ, Rafiq M, Radosa L, Hortschansky P, Cunha C, Cseresnyés Z, Krüger T, Schmidt F, Heinekamp T, Straßburger M, Löffler B, Doenst T, Lacerda JF, Campos A, Figge MT, Carvalho A, Kniemeyer O, Brakhage AA | 2023 | Aspergillus fumigatus hijacks human p11 to redirect fungal-containing phagosomes to non-degradative pathway. | Cell Host & Microbe: 31,3 | Cell Host & Microbe |
| Garloff V, Krüger T, Brakhage A, Rubio I. | 2023 | Control of TurboID-dependent biotinylation intensity in proximity ligation screens. J Proteomics. | J Proteomics 15;279:104886 | PubMed |
| Mirhakkak MH, Chen X, Ni Y, Heinekamp T, Sae-Ong T, Xu LL, Kurzai O, Barber AE, Brakhage AA, Boutin S, Schäuble S, Panagiotou G | 2023 | Genome-scale metabolic modeling of Aspergillus fumigatus strains reveals growth dependencies on the lung microbiome. | Nat Commun. 14(1):4369 | PubMed |
| Möckel L, Meusemann K, Misof B, Schwartze VU, De Fine Licht HH, Voigt K, Stielow B, de Hoog S, Beutel RG, Buellesbach J | 2022 | Phylogenetic revision and patterns of host specificity in the fungal subphylum entomophthoromycotina. | Microorganisms 10: 256 | PubMed |
| Acosta-España JD, Voigt K | 2022 | Mini Review: Risk Assessment, Clinical Manifestation, Prediction, and Prognosis of Mucormycosis: Implications for Pathogen- and Human-Derived Biomarkers. | Frontiers in Microbiology 895989 | Front. Microbiol. |
| Acosta España JD, Hassan MA, Hea-Reung P, Schwartze VU, Schäuble S, Krüger T, Kniemeyer O, Brakhage AA, Voigt K | 2022 | Novel hydrophobic binding surface proteins are instrumental for phagocytosis of Lichtheimia corymbifera by macrophages. | Medical Mycology 60 (1) :myac072P343 | Medical Mycology |
| Böttcher B, Driesch D, Krüger T, Garbe E, Gerwien F, Kniemeyer O, Brakhage AA, Vylkova S | 2022 | Impaired amino acid uptake leads to global metabolic imbalance of Candida albicans biofilms. | NPJ Biofilms Microbiomes. 8(1): 78 | PubMed |
| Montaño DE, Hartung S, Wich M, Ali R, Jungnickel B, von Lilienfeld-Toal M, Voigt K | 2022 | The TLR-NF-kB axis contributes to the monocytic inflammatory response against a virulent strain of Lichtheimia corymbifera, a causative agent of invasive mucormycosis. | Front Immunol. 13: 882921 | PubMed |
| Acosta-España JD, Voigt K | 2022 | An old confusion: Entomophthoromycosis versus mucormycosis and their main differences. | Front Microbiol. 13: 1035100. | PubMed |
| Hassan MIA, Keller M, Hillger M, Binder U, Reuter S, Herold K, Telagathoti A, Dahse HM, Wicht S, Trinks N, Nietzsche S, Deckert-Gaudig T, Deckert V, Mrowka R, Terpitz U, Saluz HP, Voigt K | 2021 | The impact of episporic modification of Lichtheimia corymbifera on virulence and interaction with phagocytes. | Comput Struct Biotechnol J 19: 880-896 | PubMed |
| Yu Y, Wolf AK, Thusek S, Heinekamp T, Bromley M, Krappmann S, Terpitz U, Voigt K, Brakhage AA, Beilhack A | 2021 | Direct visualization of fungal burden in filamentous fungus-infected silkworms. | J Fungi (Basel) 7: 136 | PubMed |
| Stanford FA, Matthies N, Cseresnyés Z, Figge MT, Hassan MIA, Voigt K | 2021 | Expression patterns in reductive iron assimilation and functional consequences during phagocytosis of Lichtheimia corymbifera, an emerging cause of Mucormycosis. | J Fungi (Basel) 7: 272 | PubMed |
| Voigt K, James TY, Kirk PM, Santiago ALCMA, Waldman B, Griffith GW, Fu M, Radek R, Strassert JFH, Wurzbacher C, Jerônimo GH, Simmons DR, Seto K, Gentekaki E, Hurdeal VG, Hyde KD, Nguyen TTT, Lee HB | 2021 | Early-diverging fungal phyla: taxonomy, species concept, ecology, distribution, anthropogenic impact, and novel phylogenetic proposals. | Fungal Divers 1-40 (Review) [Epub ahead of print] | PubMed |
| Hassan MIA, Kruse J, Krüger T, Dahse HM, Cseresnyés Z, Blango MG, Slevogt H, Hörhold F, Ast V, König R, Figge MT, Kniemeyer O, Brakhage AA, Voigt K | 2020 | Functional surface proteomic profiling reveals the host heat-shock protein A8 as a mediator of Lichtheimia corymbifera recognition by murine alveolar macrophages. | Environ Microbiol 22: 3722-40 | PubMed |
| Cseresnyés Z, Hassan MA, Dahse HD, Voigt K, Figge MT | 2020 | Quantitative impact of cell membrane fluorescence labeling on phagocytosis measurements in confrontation assays. | Front Microbiol 11: 1193 | PubMed |
| Montaño DE, Voigt K | 2020 | Host immune defense upon fungal infections with mucorales: Pathogen-immune cell interactions as drivers of inflammatory responses. | J Fungi (Basel) 6: E173 (Review) | PubMed |
| Stanford FA, Voigt K | 2020 | Iron assimilation during emerging infections caused by opportunistic fungi with emphasis on mucorales and the development of antifungal resistance. | Genes (Basel) 11: E1296 (Review) | PubMed |
| Hassan MIA, Cseresnyés Z, Al-Zaben N, Dahse HM, de Oliveira RJV, Walther G, Voigt K, Figge MT | 2019 | The geographical region of origin determines the phagocytic vulnerability of Lichtheimia strains. | Environ Microbiol 21: 4563-81 | PubMed |
| Hassan MIA, Voigt K | 2019 | Pathogenicity patterns of mucormycosis: epidemiology, interaction with immune cells and virulence factors. | Med Mycol 57: S245-S256 | PubMed |
| Gryganskyi AP, Golan J, Dolatabadi S, Mondo S, Robb S, Idnurm A, Muszewska A, Steczkiewicz K, Masonjones S, Liao HL, Gajdeczka MT, Anike F, Vuek A, Anishchenko IM, Voigt K, de Hoog GS, Smith ME, Heitman J, Vilgalys R, Stajich JE | 2018 | Phylogenetic and phylogenomic definition of Rhizopus species. | G3 (Bethesda) 8: 2007-18 | PubMed |
| Sieber P, Voigt K, Kämmer P, Brunke S, Schuster S, Linde L | 2018 | Comparative study on alternative splicing in human fungal pathogens suggests its involvement during host invasion. | Front Microbiol 9: 2313 | Front Microbiol |
| Arnesen JA, Malagocka J, Gryganskyi A, Grigoriev IV, Voigt K, Stajich JE, De Fine Licht HH | 2018 | Early diverging insect-pathogenic fungi of the order Entomophthorales possess diverse and unique subtilisin-like serine proteases. | G3 (Bethesda) 8: 3311-19 | PubMed |
| Schulze B, Rambach G, Schwartze VU, Voigt K, Schubert K, Speth C, Jacobsen ID | 2017 | Ketoacidosis alone does not predispose to mucormycosis by Lichtheimia in a murine pulmonary infection model. | Virulence 8: 1657-67 | PubMed |